m6A-centered Drug Response Information
General Information of the Drug (ID: M6ADRUG0044)
| Name |
Rapamycin
|
||||
|---|---|---|---|---|---|
| Synonyms |
Rapamune; Rapamycin (Sirolimus); AY-22989; Rapammune; sirolimusum; WY-090217; RAPA; Antibiotic AY 22989; AY 22989; UNII-W36ZG6FT64; CCRIS 9024; CHEBI:9168; SILA 9268A; W36ZG6FT64; HSDB 7284; C51H79NO13; NSC 226080; DE-109; NCGC00021305-05; DSSTox_CID_3582; DSSTox_RID_77091; DSSTox_GSID_23582; Cypher; Supralimus; Wy 090217; Perceiva; RAP; RPM; Rapamycin from Streptomyces hygroscopicus; SIIA 9268A; LCP-Siro; MS-R001; Rapamune (TN); Rapamycin (TN); Sirolimus (RAPAMUNE); Rapamycin C-7, analog 4; Sirolimus (USAN/INN); Sirolimus [USAN:BAN:INN]; Sirolimus, Rapamune,Rapamycin; Heptadecahydro-9,27-dihydroxy-3-[(1R)-2-[(1S,3R,4R)-4-hydroxy; 23,27-Epoxy-3H-pyrido(2,1-c)(1,4)oxaazacyclohentriacontine; 23,27-Epoxy-3H-pyrido[2,1-c][1,4]oxaazacyclohentriacontine; 23,27-epoxy-3H-pyrido[2,1-c][1,4]oxaazacyclohentriacontine-1,5,11,28,29; 3H-pyrido(2,1-c)(1,4)oxaazacyclohentriacontine-1,5,11,28,29(4H,6H,31H)-pentone; Sirolimus (MTOR inhibitor)
Click to Show/Hide
|
||||
| Status | Approved | [1] | |||
| Structure |
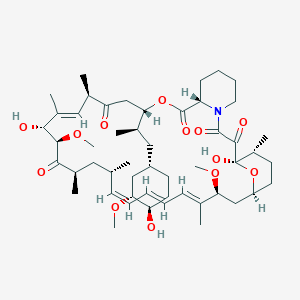 |
||||
|
3D MOL
|
|||||
| Formula |
C51H79NO13
|
||||
| InChI |
InChI=1S/C51H79NO13/c1-30-16-12-11-13-17-31(2)42(61-8)28-38-21-19-36(7)51(60,65-38)48(57)49(58)52-23-15-14-18-39(52)50(59)64-43(33(4)26-37-20-22-40(53)44(27-37)62-9)29-41(54)32(3)25-35(6)46(56)47(63-10)45(55)34(5)24-30/h11-13,16-17,25,30,32-34,36-40,42-44,46-47,53,56,60H,14-15,18-24,26-29H2,1-10H3/b13-11+,16-12+,31-17+,35-25+/t30-,32-,33-,34-,36-,37+,38+,39+,40-,42+,43+,44-,46-,47+,51-/m1/s1
|
||||
| InChIKey |
QFJCIRLUMZQUOT-HPLJOQBZSA-N
|
||||
| PubChem CID | |||||
| TTD Drug ID | |||||
| DrugBank ID | |||||
Full List of m6A Targets Related to This Drug
Cyclic AMP-dependent transcription factor ATF-4 (ATF4)
| In total 2 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [2] | |||
| Response Summary | In colorectal cancer, Glutaminolysis inhibition upregulated ATF4 expression in an m6A-dependent manner to activate pro-survival autophagy through transcriptional activation of the mTOR inhibitor DDIT4. Determined the relationship between FTO alpha-ketoglutarate dependent dioxygenase (FTO), YTH N6-methyladenosine RNA binding protein 2 (YTHDF2), and ATF4. Cyclic AMP-dependent transcription factor ATF-4 (ATF4) transcriptionally upregulated DDIT4 to suppress mTOR, which induced pro-survival autophagy during glutaminolysis inhibition. | |||
| Responsed Disease | Colorectal cancer | ICD-11: 2B91 | ||
| Target Regulator | YTH domain-containing family protein 2 (YTHDF2) | READER | ||
| Target Regulation | Down regulation | |||
| Pathway Response | mTOR signaling pathway | hsa04150 | ||
| Cell Process | RNA decay | |||
| Cell growth and death | ||||
| Cell autophagy | ||||
| In-vitro Model | HCT 116 | Colon carcinoma | Homo sapiens | CVCL_0291 |
| SW480 | Colon adenocarcinoma | Homo sapiens | CVCL_0546 | |
| Experiment 2 Reporting the m6A-centered Drug Response by This Target Gene | [2] | |||
| Response Summary | In colorectal cancer, Glutaminolysis inhibition upregulated ATF4 expression in an m6A-dependent manner to activate pro-survival autophagy through transcriptional activation of the mTOR inhibitor DDIT4. Determined the relationship between FTO alpha-ketoglutarate dependent dioxygenase (FTO), YTH N6-methyladenosine RNA binding protein 2 (YTHDF2), and ATF4. Cyclic AMP-dependent transcription factor ATF-4 (ATF4) transcriptionally upregulated DDIT4 to suppress mTOR, which induced pro-survival autophagy during glutaminolysis inhibition. | |||
| Responsed Disease | Colorectal cancer | ICD-11: 2B91 | ||
| Target Regulator | Fat mass and obesity-associated protein (FTO) | ERASER | ||
| Pathway Response | mTOR signaling pathway | hsa04150 | ||
| Cell Process | RNA decay | |||
| Cell growth and death | ||||
| Cell autophagy | ||||
| In-vitro Model | HCT 116 | Colon carcinoma | Homo sapiens | CVCL_0291 |
| SW480 | Colon adenocarcinoma | Homo sapiens | CVCL_0546 | |
eIF4E-binding protein 1 (4EBP1/EIF4EBP1)
| In total 1 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [3] | |||
| Response Summary | METTL3 promotes the progression of retinoblastoma through PI3K/AKT/mTOR pathways in vitro and in vivo. METTL3 has an impact on the PI3K-AKT-mTOR-P70S6K/eIF4E-binding protein 1 (4EBP1/EIF4EBP1) pathway. The cell proliferation results show that the stimulatory function of METTL3 is lost after rapamycin treatment. | |||
| Responsed Disease | Retinoblastoma | ICD-11: 2D02.2 | ||
| Target Regulator | Methyltransferase-like 3 (METTL3) | WRITER | ||
| Target Regulation | Down regulation | |||
| Pathway Response | PI3K-Akt signaling pathway | hsa04151 | ||
| mTOR signaling pathway | hsa04150 | |||
| Cell Process | Cell proliferation | |||
| Cell migration | ||||
| Cell invasion | ||||
| Cell apoptosis | ||||
| In-vitro Model | WERI-Rb-1 | Retinoblastoma | Homo sapiens | CVCL_1792 |
| Y-79 | Retinoblastoma | Homo sapiens | CVCL_1893 | |
| In-vivo Model | To establish a subcutaneous tumour model in nude mice, 2 × 107 Y79 cells (METTL3 knockdown group: shNC, shRNA1 and shRNA2; METTL3 up-regulated group: NC and METLL3) were resuspended in 1 mL of pre-cooled PBS, and 200 uL of the cell suspension was injected subcutaneously into the left side of the armpit to investigate tumour growth (4 × 106 per mouse). | |||
Eukaryotic translation initiation factor 4 gamma 1 (EIF4G1)
| In total 2 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [4] | |||
| Response Summary | Rapamycin inhibited FTO activity, and directly targeted Eukaryotic translation initiation factor 4 gamma 1 (EIF4G1) transcripts and mediated their expression in an m6A-dependent manner in oral squamous cell carcinoma. After FTO silencing, YTHDF2 captured eIF4G1 transcripts containing m6A, resulting in mRNA degradation and decreased expression of eIF4G1 protein, thereby promoting autophagy and reducing tumor occurrence. | |||
| Responsed Disease | Oral squamous cell carcinoma | ICD-11: 2B6E.0 | ||
| Target Regulator | Fat mass and obesity-associated protein (FTO) | ERASER | ||
| Target Regulation | Up regulation | |||
| Pathway Response | Autophagy | hsa04140 | ||
| Cell Process | Cell autophagy | |||
| Experiment 2 Reporting the m6A-centered Drug Response by This Target Gene | [4] | |||
| Response Summary | Rapamycin inhibited FTO activity, and directly targeted Eukaryotic translation initiation factor 4 gamma 1 (EIF4G1) transcripts and mediated their expression in an m6A-dependent manner in oral squamous cell carcinoma. After FTO silencing, YTHDF2 captured eIF4G1 transcripts containing m6A, resulting in mRNA degradation and decreased expression of eIF4G1 protein, thereby promoting autophagy and reducing tumor occurrence. | |||
| Responsed Disease | Oral squamous cell carcinoma | ICD-11: 2B6E.0 | ||
| Target Regulator | YTH domain-containing family protein 2 (YTHDF2) | READER | ||
| Target Regulation | Down regulation | |||
| Pathway Response | Autophagy | hsa04140 | ||
| Cell Process | Cell autophagy | |||
Mammalian target of rapamycin complex 1 (mTORC1)
| In total 1 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [5] | |||
| Response Summary | Mammalian target of rapamycin complex 1 (mTORC1) pathway is highly activated in rbm15-deficient hepatocytes. Rapamycin treatment partially restored normal hepatic gene expression as well as the nuclear location of the transcription factor Hnf4a. Taken together, these results reveal an unexpected role of Rbm15 in liver maturation. | |||
| Responsed Disease | Liver disease | ICD-11: DB9Z | ||
| Target Regulator | RNA-binding motif protein 15 (RBM15) | WRITER | ||
| Target Regulation | Down regulation | |||
| Pathway Response | mTOR signaling pathway | hsa04150 | ||
| Cell Process | Cell proliferation and apoptosis | |||
| In-vivo Model | Zebrafish (Danio rerio) AB strain-derived Tg(lfabp:Dendra2-NTR)cq1 was used as WT, and rbm15cq96 mutant was generated by ENU treatment. | |||
Phosphatidylinositol 3-kinase regulatory subunit beta (PI3K-p85/PIK3R2)
| In total 1 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [3] | |||
| Response Summary | METTL3 promotes the progression of retinoblastoma through Phosphatidylinositol 3-kinase regulatory subunit beta (PI3K-p85/PIK3R2)/AKT/mTOR pathways in vitro and in vivo. METTL3 has an impact on the PI3K-AKT-mTOR-P70S6K/4EBP1 pathway. The cell proliferation results show that the stimulatory function of METTL3 is lost after rapamycin treatment. | |||
| Responsed Disease | Retinoblastoma | ICD-11: 2D02.2 | ||
| Target Regulator | Methyltransferase-like 3 (METTL3) | WRITER | ||
| Target Regulation | Up regulation | |||
| Pathway Response | PI3K-Akt signaling pathway | hsa04151 | ||
| mTOR signaling pathway | hsa04150 | |||
| Apoptosis | hsa04210 | |||
| Cell Process | Cell proliferation | |||
| Cell migration | ||||
| Cell invasion | ||||
| Cell apoptosis | ||||
| In-vitro Model | WERI-Rb-1 | Retinoblastoma | Homo sapiens | CVCL_1792 |
| Y-79 | Retinoblastoma | Homo sapiens | CVCL_1893 | |
| In-vivo Model | To establish a subcutaneous tumour model in nude mice, 2 × 107 Y79 cells (METTL3 knockdown group: shNC, shRNA1 and shRNA2; METTL3 up-regulated group: NC and METLL3) were resuspended in 1 mL of pre-cooled PBS, and 200 uL of the cell suspension was injected subcutaneously into the left side of the armpit to investigate tumour growth (4 × 106 per mouse). | |||
RAC-alpha serine/threonine-protein kinase (AKT1)
| In total 1 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [3] | |||
| Response Summary | METTL3 promotes the progression of retinoblastoma through PI3K/RAC-alpha serine/threonine-protein kinase (AKT1)/mTOR pathways in vitro and in vivo. METTL3 has an impact on the PI3K-AKT-mTOR-P70S6K/4EBP1 pathway. The cell proliferation results show that the stimulatory function of METTL3 is lost after rapamycin treatment. | |||
| Responsed Disease | Retinoblastoma | ICD-11: 2D02.2 | ||
| Target Regulator | Methyltransferase-like 3 (METTL3) | WRITER | ||
| Target Regulation | Up regulation | |||
| Pathway Response | PI3K-Akt signaling pathway | hsa04151 | ||
| mTOR signaling pathway | hsa04150 | |||
| Apoptosis | hsa04210 | |||
| Cell Process | Cell proliferation | |||
| Cell migration | ||||
| Cell invasion | ||||
| Cell apoptosis | ||||
| In-vitro Model | WERI-Rb-1 | Retinoblastoma | Homo sapiens | CVCL_1792 |
| Y-79 | Retinoblastoma | Homo sapiens | CVCL_1893 | |
| In-vivo Model | To establish a subcutaneous tumour model in nude mice, 2 × 107 Y79 cells (METTL3 knockdown group: shNC, shRNA1 and shRNA2; METTL3 up-regulated group: NC and METLL3) were resuspended in 1 mL of pre-cooled PBS, and 200 uL of the cell suspension was injected subcutaneously into the left side of the armpit to investigate tumour growth (4 × 106 per mouse). | |||
Ribosomal protein S6 kinase beta-1 (RPS6KB1/p70S6K)
| In total 1 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [3] | |||
| Response Summary | METTL3 promotes the progression of retinoblastoma through PI3K/AKT/mTOR pathways in vitro and in vivo. METTL3 has an impact on the PI3K-AKT-mTOR-Ribosomal protein S6 kinase beta-1 (RPS6KB1/p70S6K)/4EBP1 pathway. The cell proliferation results show that the stimulatory function of METTL3 is lost after rapamycin treatment. | |||
| Responsed Disease | Retinoblastoma | ICD-11: 2D02.2 | ||
| Target Regulator | Methyltransferase-like 3 (METTL3) | WRITER | ||
| Target Regulation | Up regulation | |||
| Pathway Response | PI3K-Akt signaling pathway | hsa04151 | ||
| mTOR signaling pathway | hsa04150 | |||
| Cell Process | Cell proliferation | |||
| Cell migration | ||||
| Cell invasion | ||||
| Cell apoptosis | ||||
| In-vitro Model | WERI-Rb-1 | Retinoblastoma | Homo sapiens | CVCL_1792 |
| Y-79 | Retinoblastoma | Homo sapiens | CVCL_1893 | |
| In-vivo Model | To establish a subcutaneous tumour model in nude mice, 2 × 107 Y79 cells (METTL3 knockdown group: shNC, shRNA1 and shRNA2; METTL3 up-regulated group: NC and METLL3) were resuspended in 1 mL of pre-cooled PBS, and 200 uL of the cell suspension was injected subcutaneously into the left side of the armpit to investigate tumour growth (4 × 106 per mouse). | |||
Sequestosome-1 (SQSTM1)
| In total 1 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [6] | |||
| Response Summary | The m6A changes caused by FTO influence the stability of ULK1 transcripts, likely through a YTHDF2-dependent manner.Under both basal and rapamycin-induced autophagy conditions, depletion of FTO significantly reduced the formation of GFP-LC3B puncta. The level of Sequestosome-1 (SQSTM1)/SQSTM1 (an autophagy substrate) was higher in FTO-knockdown cells than that in control cells. FTO specifically upregulates the ULK1 protein abundance. ULK1 mRNA undergoes m6A modification in the 3'-UTR and the m6A-marked ULK1 transcripts can further be targeted for degradation by YTHDF2. | |||
| Target Regulator | Fat mass and obesity-associated protein (FTO) | ERASER | ||
| Target Regulation | Down regulation | |||
| Pathway Response | Autophagy | hsa04140 | ||
| Cell Process | RNA stability | |||
| Cell autophagy | ||||
| In-vitro Model | HEK293T | Normal | Homo sapiens | CVCL_0063 |
| HeLa | Endocervical adenocarcinoma | Homo sapiens | CVCL_0030 | |
Serine/threonine-protein kinase mTOR (MTOR)
| In total 3 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [2] | |||
| Response Summary | In colorectal cancer, Glutaminolysis inhibition upregulated ATF4 expression in an m6A-dependent manner to activate pro-survival autophagy through transcriptional activation of the mTOR inhibitor DDIT4. Determined the relationship between FTO alpha-ketoglutarate dependent dioxygenase (FTO), YTH N6-methyladenosine RNA binding protein 2 (YTHDF2), and ATF4. Serine/threonine-protein kinase mTOR (MTOR) transcriptionally upregulated DDIT4 to suppress mTOR, which induced pro-survival autophagy during glutaminolysis inhibition. | |||
| Responsed Disease | Colorectal cancer | ICD-11: 2B91 | ||
| Target Regulator | YTH domain-containing family protein 2 (YTHDF2) | READER | ||
| Target Regulation | Up regulation | |||
| Pathway Response | mTOR signaling pathway | hsa04150 | ||
| Autophagy | hsa04140 | |||
| Cell Process | RNA decay | |||
| Cell growth and death | ||||
| Cell autophagy | ||||
| In-vitro Model | HCT 116 | Colon carcinoma | Homo sapiens | CVCL_0291 |
| SW480 | Colon adenocarcinoma | Homo sapiens | CVCL_0546 | |
| Experiment 2 Reporting the m6A-centered Drug Response by This Target Gene | [2] | |||
| Response Summary | In colorectal cancer, Glutaminolysis inhibition upregulated ATF4 expression in an m6A-dependent manner to activate pro-survival autophagy through transcriptional activation of the mTOR inhibitor DDIT4. Determined the relationship between FTO alpha-ketoglutarate dependent dioxygenase (FTO), YTH N6-methyladenosine RNA binding protein 2 (YTHDF2), and ATF4. Serine/threonine-protein kinase mTOR (MTOR) transcriptionally upregulated DDIT4 to suppress mTOR, which induced pro-survival autophagy during glutaminolysis inhibition. | |||
| Responsed Disease | Colorectal cancer | ICD-11: 2B91 | ||
| Target Regulator | Fat mass and obesity-associated protein (FTO) | ERASER | ||
| Pathway Response | mTOR signaling pathway | hsa04150 | ||
| Autophagy | hsa04140 | |||
| Cell Process | RNA decay | |||
| Cell growth and death | ||||
| Cell autophagy | ||||
| In-vitro Model | HCT 116 | Colon carcinoma | Homo sapiens | CVCL_0291 |
| SW480 | Colon adenocarcinoma | Homo sapiens | CVCL_0546 | |
| Experiment 3 Reporting the m6A-centered Drug Response by This Target Gene | [3] | |||
| Response Summary | METTL3 promotes the progression of retinoblastoma through PI3K/AKT/Serine/threonine-protein kinase mTOR (MTOR) pathways in vitro and in vivo. METTL3 has an impact on the PI3K-AKT-mTOR-P70S6K/4EBP1 pathway. The cell proliferation results show that the stimulatory function of METTL3 is lost after rapamycin treatment. | |||
| Responsed Disease | Retinoblastoma | ICD-11: 2D02.2 | ||
| Target Regulator | Methyltransferase-like 3 (METTL3) | WRITER | ||
| Target Regulation | Up regulation | |||
| Pathway Response | PI3K-Akt signaling pathway | hsa04151 | ||
| mTOR signaling pathway | hsa04150 | |||
| Cell Process | Cell proliferation | |||
| Cell migration | ||||
| Cell invasion | ||||
| Cell apoptosis | ||||
| In-vitro Model | WERI-Rb-1 | Retinoblastoma | Homo sapiens | CVCL_1792 |
| Y-79 | Retinoblastoma | Homo sapiens | CVCL_1893 | |
| In-vivo Model | To establish a subcutaneous tumour model in nude mice, 2 × 107 Y79 cells (METTL3 knockdown group: shNC, shRNA1 and shRNA2; METTL3 up-regulated group: NC and METLL3) were resuspended in 1 mL of pre-cooled PBS, and 200 uL of the cell suspension was injected subcutaneously into the left side of the armpit to investigate tumour growth (4 × 106 per mouse). | |||
Serine/threonine-protein kinase ULK1 (ULK1/ATG1)
| In total 2 item(s) under this target gene | ||||
| Experiment 1 Reporting the m6A-centered Drug Response by This Target Gene | [6] | |||
| Response Summary | The m6A changes caused by FTO influence the stability of ULK1 transcripts, likely through a YTHDF2-dependent manner.Under both basal and rapamycin-induced autophagy conditions, depletion of FTO significantly reduced the formation of GFP-LC3B puncta. The level of p62/SQSTM1 (an autophagy substrate) was higher in FTO-knockdown cells than that in control cells. FTO specifically upregulates the Serine/threonine-protein kinase ULK1 (ULK1) protein abundance. ULK1 mRNA undergoes m6A modification in the 3'-UTR and the m6A-marked ULK1 transcripts can further be targeted for degradation by YTHDF2. | |||
| Target Regulator | Fat mass and obesity-associated protein (FTO) | ERASER | ||
| Target Regulation | Up regulation | |||
| Pathway Response | Autophagy | hsa04140 | ||
| Cell Process | RNA stability | |||
| Cell autophagy | ||||
| In-vitro Model | HEK293T | Normal | Homo sapiens | CVCL_0063 |
| HeLa | Endocervical adenocarcinoma | Homo sapiens | CVCL_0030 | |
| Experiment 2 Reporting the m6A-centered Drug Response by This Target Gene | [6] | |||
| Response Summary | The m6A changes caused by FTO influence the stability of ULK1 transcripts, likely through a YTHDF2-dependent manner.Under both basal and rapamycin-induced autophagy conditions, depletion of FTO significantly reduced the formation of GFP-LC3B puncta. The level of p62/SQSTM1 (an autophagy substrate) was higher in FTO-knockdown cells than that in control cells. FTO specifically upregulates the Serine/threonine-protein kinase ULK1 (ULK1) protein abundance. ULK1 mRNA undergoes m6A modification in the 3'-UTR and the m6A-marked ULK1 transcripts can further be targeted for degradation by YTHDF2. | |||
| Target Regulator | YTH domain-containing family protein 2 (YTHDF2) | READER | ||
| Target Regulation | Up regulation | |||
| Pathway Response | Autophagy | hsa04140 | ||
| Cell Process | RNA stability | |||
| Cell autophagy | ||||
| In-vitro Model | HEK293T | Normal | Homo sapiens | CVCL_0063 |
| HeLa | Endocervical adenocarcinoma | Homo sapiens | CVCL_0030 | |
Full List of Crosstalk(s) between m6A Modification and Epigenetic Regulation Related to This Drug
| In total 7 item(s) under this drug | ||
| Crosstalk ID: M6ACROT02224 | ||
| m6A Regulator | Methyltransferase-like 14 (METTL14) | |
| m6A Target | Transmembrane protein 127 (TMEM127) | |
| Epigenetic Regulator | DNA (cytosine-5)-methyltransferase 3B (DNMT3B) | |
| Regulated Target | Methyltransferase-like protein 14 (METTL14) | |
| Crosstalk relationship | DNA modification → m6A | |
| Disease | Breast cancer | |
| Crosstalk ID: M6ACROT02248 | ||
| m6A Regulator | Methyltransferase-like 14 (METTL14) | |
| m6A Target | Transmembrane protein 127 (TMEM127) | |
| Epigenetic Regulator | Cysteine methyltransferase DNMT3A (DNMT3A) | |
| Regulated Target | Methyltransferase-like protein 14 (METTL14) | |
| Crosstalk relationship | DNA modification → m6A | |
| Disease | Breast cancer | |
| Crosstalk ID: M6ACROT02272 | ||
| m6A Regulator | Methyltransferase-like 14 (METTL14) | |
| m6A Target | Transmembrane protein 127 (TMEM127) | |
| Epigenetic Regulator | DNA (cytosine-5)-methyltransferase 1 (DNMT1) | |
| Regulated Target | Methyltransferase-like protein 14 (METTL14) | |
| Crosstalk relationship | DNA modification → m6A | |
| Disease | Breast cancer | |
| Crosstalk ID: M6ACROT03334 | ||
| m6A Regulator | YTH domain-containing family protein 2 (YTHDF2) | |
| m6A Target | Cyclic AMP-dependent transcription factor ATF-4 (ATF4) | |
| Epigenetic Regulator | Histone acetyltransferase p300 (P300) | |
| Regulated Target | Histone H3 lysine 18 lactylation (H3K18la) | |
| Crosstalk relationship | Histone modification → m6A | |
| Disease | Colorectal cancer | |
| Crosstalk ID: M6ACROT03340 | ||
| m6A Regulator | YTH domain-containing family protein 2 (YTHDF2) | |
| m6A Target | Serine/threonine-protein kinase mTOR (MTOR) | |
| Epigenetic Regulator | Histone acetyltransferase p300 (P300) | |
| Regulated Target | Histone H3 lysine 18 lactylation (H3K18la) | |
| Crosstalk relationship | Histone modification → m6A | |
| Disease | Colorectal cancer | |
| Crosstalk ID: M6ACROT03432 | ||
| m6A Regulator | Fat mass and obesity-associated protein (FTO) | |
| m6A Target | Cyclic AMP-dependent transcription factor ATF-4 (ATF4) | |
| Epigenetic Regulator | Histone deacetylase 1 (HDAC1) | |
| Regulated Target | Histone H3 lysine 27 acetylation (H3K27ac) | |
| Crosstalk relationship | Histone modification → m6A | |
| Disease | Colorectal cancer | |
| Crosstalk ID: M6ACROT03438 | ||
| m6A Regulator | Fat mass and obesity-associated protein (FTO) | |
| m6A Target | Serine/threonine-protein kinase mTOR (MTOR) | |
| Epigenetic Regulator | Histone deacetylase 1 (HDAC1) | |
| Regulated Target | Histone H3 lysine 27 acetylation (H3K27ac) | |
| Crosstalk relationship | Histone modification → m6A | |
| Disease | Colorectal cancer | |
References